| Gene | Hpcal4 |
| Chromosome | 4 |
| Deletion Start | 123,188,707 |
| Deletion End | 123,190,878 |
| Deletion Size | 2,172 |
| Genome Build | 38 |
|
|
Primer Genomic Coordinates on Chromosome 4
Please be aware that these oligonucleotides have been validated only by Taqman assay.
| Primer | Start | End | Sequence |
| TUF | 123,189,076 | 123,189,095 | TTCGACAAGAACGGCGATGG |
| TUP | 123,189,098 | 123,189,123 | CCATCGACTTCCGGGAGTTCATCTGC |
| TUR | 123,189,126 | 123,189,145 | CACGGGAGGTGACAGAAAGG |
| TDF | 123,189,903 | 123,189,923 | CACGGTGGCTCACTTCTGTAG |
| TDP | 123,189,924 | 123,189,948 | TCCCAACACTCAGGAAGCTAAGGCA |
| TDR | 123,189,959 | 123,189,979 | GCTCAGCCTGTCCTTGAACTC |
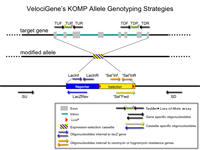
PCR Primers
| Primer | Sequence |
| SU | AGAGCTCGCTGAATGGGTG |
| SD | GGTGGAGCTAGTGACAGAC |
| LacInR | TTGACTGTAGCGGCTGATGTTG |
| LacInf | GGTAAACTGGCTCGGATTAGGG |
| NeoInR | TACTTTCTCGGCAGGAGCAAGGTG |
| NeoInF | TTCGGCTATGACTGGGCACAACAG |
| LacInZRev | GTCTGTCCTAGCTTCCTCACTG |
| NeoFwd | TCATTCTCAGTATTGTTTTGCC |
Predicted PCR Products
| Forward Primer | Reverse Primer | Length | WT | HET | KO |
| SU | LacZRev | 307 bp | - | + | + |
| TUF | TUR | 70 bp | + | + | - |
| TDF | TDR | 77 bp | + | + | - |
| NeoFwd | SD | 302 bp | - | + | + |
| LacInf | LacInR | 210 bp | - | + | + |
| NeoInf | NeoInR | 282 bp | - | + | + |
Available Clones
ES Cell ordering information can be found at the
KOMP Repository.
| Clone Name | Parental ES Cell | Mouse Strain | Cassette Copy Number |
| 10300A-A2 | VGB6 | C57BL/6NTac | |
| 10300A-A4 | VGB6 | C57BL/6NTac | |
| 10300A-E2 | VGB6 | C57BL/6NTac | |
| 10300A-E6 | VGB6 | C57BL/6NTac | |
| 10300B-B10 | VGB6 | C57BL/6NTac | |
| 10300B-C6 | VGB6 | C57BL/6NTac | |