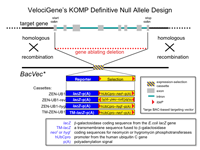
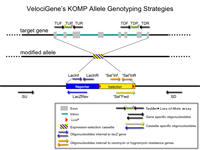
16145 Arhgap36 Status: ES cell colonies screened / QC no positives
| Gene | Arhgap36 |
| Chromosome | X |
| Deletion Start | 49,499,152 |
| Deletion End | 49,470,610 |
| Deletion Size | 28,543 |
| Genome Build | 38 |
|
|
Primer Genomic Coordinates on Chromosome X
Please be aware that these oligonucleotides have been validated only by Taqman assay.
| Primer | Start | End | Sequence |
| TUF | 49,471,204 | 49,471,223 | CATGTAGTTGGCAGGCAGTC |
| TUP | 49,471,224 | 49,471,249 | TGGGACAGCTTAGGCTCTGCTAACTA |
| TUR | 49,471,256 | 49,471,274 | TGAGCAGTTCAGGGCAATG |
| TDF | 49,498,567 | 49,498,589 | TCTTGCAGAGGTTCCCTATCTTG |
| TDP | 49,498,599 | 49,498,626 | CAAACAACGAAGATACCGACTCAGACGC |
| TDR | 49,498,631 | 49,498,655 | GGGCTTCATCATTACCTGGGATTAC |
PCR Primers
| Primer | Sequence |
| SU | TGCTCTAGACAGGAGCTGGG |
| SD | TTTCCAGTTGAGCTCTTGGG |
| LacInR | TTGACTGTAGCGGCTGATGTTG |
| LacInf | GGTAAACTGGCTCGGATTAGGG |
| NeoInR | TACTTTCTCGGCAGGAGCAAGGTG |
| NeoInF | TTCGGCTATGACTGGGCACAACAG |
| LacInZRev | GTCTGTCCTAGCTTCCTCACTG |
| NeoFwd | TCATTCTCAGTATTGTTTTGCC |
Predicted PCR Products
| Forward Primer | Reverse Primer | Length | WT | HET | KO |
| SU | LacZRev | bp | - | + | + |
| TUF | TUR | 71 bp | + | + | - |
| TDF | TDR | 89 bp | + | + | - |
| NeoFwd | SD | bp | - | + | + |
| LacInf | LacInR | 210 bp | - | + | + |
| NeoInf | NeoInR | 282 bp | - | + | + |